Note
Go to the end to download the full example code.
Filtering Data (Averaging Neighboring Patterns etc.)#
If you have a low number of counts in your data, you may want to filter the data to remove noise. This can be done using the filter function which applies some function to the entire dataset and returns a filtered dataset of the same shape.
In this example, we will use the filter function to apply a Gaussian filter in only the real space dimensions of the dataset. This is useful for removing noise as it acts like a spatial bandpass filter for high frequencies (noise) in the dataset.
Because the STEM probe is Gaussian-like, and the convolution of two Gaussian functions is another Gaussian function, the Gaussian filter is a good choice for filtering and is equivalent to aquiring the data with a larger probe size equal to \(\sigma_{Filtererd} = \sqrt{\sigma_{Beam}^2 + \sigma_{Filter}^2}\). The benefit is that the total aquisition time is much shorter than if the probe size was increased to reach the same number of counts. For detectors with low saturation counts, this can be a significant advantage.
from scipy.ndimage import gaussian_filter
from dask_image.ndfilters import gaussian_filter as dask_gaussian_filter
import pyxem as pxm
import hyperspy.api as hs
import numpy as np
s = pxm.data.tilt_boundary_data()
s_filtered = s.filter(
gaussian_filter, sigma=1.0, inplace=False
) # Gaussian filter with sigma=1.0
s_filtered2 = s.filter(
gaussian_filter, sigma=(1.0, 1.0, 0, 0), inplace=False
) # Only filter in real space
hs.plot.plot_images(
[s.inav[3, 3], s_filtered.inav[3, 3], s_filtered2.inav[3, 3]],
label=["Original", "GaussFilt(all)", "GaussFilt(real space)"],
tight_layout=True,
vmax="99th",
)
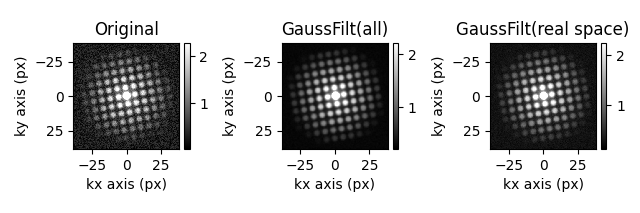
[<Axes: title={'center': 'Original'}, xlabel='kx axis (px)', ylabel='ky axis (px)'>, <Axes: title={'center': 'GaussFilt(all)'}, xlabel='kx axis (px)', ylabel='ky axis (px)'>, <Axes: title={'center': 'GaussFilt(real space)'}, xlabel='kx axis (px)', ylabel='ky axis (px)'>]
The filter function can also be used with a custom function as long as the function takes a numpy array as input and returns a numpy array of the same shape.
def custom_filter(array):
filtered = gaussian_filter(array, sigma=1.0)
return filtered - np.mean(filtered)
s_filtered3 = s.filter(custom_filter, inplace=False) # Custom filter
hs.plot.plot_images(
[s.inav[3, 3], s_filtered3.inav[3, 3]],
label=["Original", "GaussFilt(Custom)"],
tight_layout=True,
vmax="99th",
)
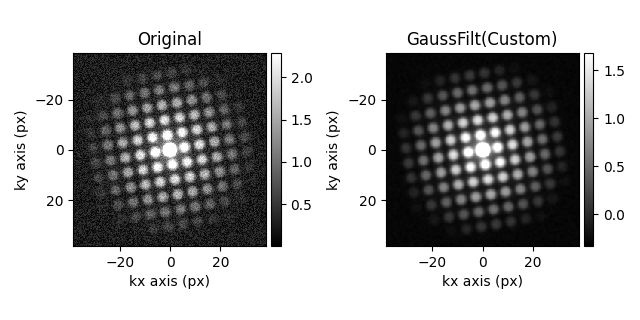
[<Axes: title={'center': 'Original'}, xlabel='kx axis (px)', ylabel='ky axis (px)'>, <Axes: title={'center': 'GaussFilt(Custom)'}, xlabel='kx axis (px)', ylabel='ky axis (px)'>]
For lazy datasets, functions which operate on dask arrays can be used. For example, the gaussian_filter function from scipy.ndimage is replaced with the dask_image version which operates on dask arrays.
s = s.as_lazy() # Convert to lazy dataset
s_filtered4 = s.filter(
dask_gaussian_filter, sigma=1.0, inplace=False
) # Gaussian filter with sigma=1.0
hs.plot.plot_images(
[s_filtered.inav[3, 3], s_filtered4.inav[3, 3]],
label=["GaussFilt", "GaussFilt(Lazy)"],
tight_layout=True,
vmax="99th",
)
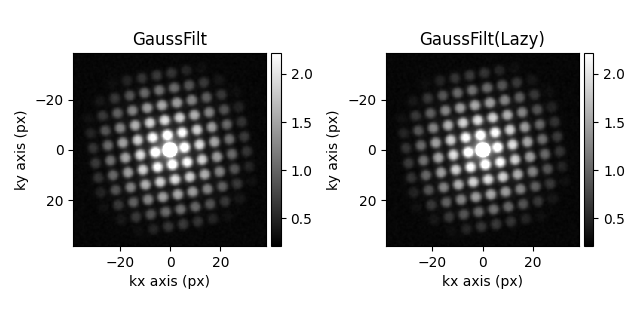
[<Axes: title={'center': 'GaussFilt'}, xlabel='kx axis (px)', ylabel='ky axis (px)'>, <Axes: title={'center': 'GaussFilt(Lazy)'}, xlabel='kx axis (px)', ylabel='ky axis (px)'>]
Total running time of the script: (0 minutes 1.315 seconds)
