Note
Go to the end to download the full example code.
Background subtraction#
If your diffraction data is noisy, you might want to subtract the background from the dataset. Pyxem offers some built-in functionality for this, with the subtract_diffraction_background class method. Custom filtering is also possible, an example is shown in the ‘Filtering Data’-example.
import pyxem as pxm
import hyperspy.api as hs
s = pxm.data.tilt_boundary_data()
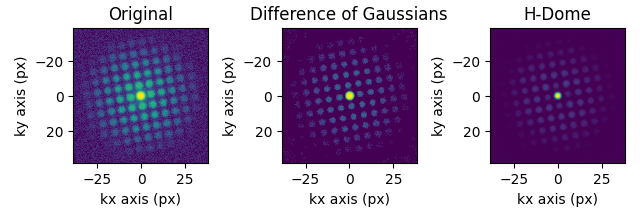
s_filtered = s.subtract_diffraction_background(
"difference of gaussians",
inplace=False,
min_sigma=3,
max_sigma=20,
)
s_filtered_h = s.subtract_diffraction_background("h-dome", inplace=False, h=0.7)
hs.plot.plot_images(
[s.inav[2, 2], s_filtered.inav[2, 2], s_filtered_h.inav[2, 2]],
label=["Original", "Difference of Gaussians", "H-Dome"],
tight_layout=True,
norm="symlog",
cmap="viridis",
colorbar=None,
)
0%| | 0/33 [00:00<?, ?it/s]
33%|███▎ | 11/33 [00:00<00:00, 89.02it/s]
61%|██████ | 20/33 [00:00<00:00, 86.67it/s]
88%|████████▊ | 29/33 [00:00<00:00, 73.65it/s]
100%|██████████| 33/33 [00:00<00:00, 73.78it/s]
/home/docs/checkouts/readthedocs.org/user_builds/pyxem/envs/1139/lib/python3.11/site-packages/pyxem/utils/_background_subtraction.py:99: FutureWarning: `square` is deprecated since version 0.25 and will be removed in version 0.27. Use `skimage.morphology.footprint_rectangle` instead.
regional_filter(frame / max_value, **kwargs), footprint=square(3)
/home/docs/checkouts/readthedocs.org/user_builds/pyxem/envs/1139/lib/python3.11/site-packages/hyperspy/misc/utils.py:1439: UserWarning: Possible precision loss converting image of type float32 to uint8 as required by rank filters. Convert manually using skimage.util.img_as_ubyte to silence this warning.
output = function(test_data, **kwargs)
0%| | 0/33 [00:00<?, ?it/s]/home/docs/checkouts/readthedocs.org/user_builds/pyxem/envs/1139/lib/python3.11/site-packages/pyxem/utils/_background_subtraction.py:99: FutureWarning: `square` is deprecated since version 0.25 and will be removed in version 0.27. Use `skimage.morphology.footprint_rectangle` instead.
regional_filter(frame / max_value, **kwargs), footprint=square(3)
/home/docs/checkouts/readthedocs.org/user_builds/pyxem/envs/1139/lib/python3.11/site-packages/hyperspy/misc/utils.py:1362: UserWarning: Possible precision loss converting image of type float32 to uint8 as required by rank filters. Convert manually using skimage.util.img_as_ubyte to silence this warning.
output_array[islice] = function(data[islice], **kwargs)
15%|█▌ | 5/33 [00:00<00:00, 44.24it/s]
33%|███▎ | 11/33 [00:00<00:00, 25.41it/s]
45%|████▌ | 15/33 [00:00<00:00, 28.45it/s]
58%|█████▊ | 19/33 [00:00<00:00, 22.71it/s]
67%|██████▋ | 22/33 [00:00<00:00, 22.36it/s]
76%|███████▌ | 25/33 [00:01<00:00, 23.70it/s]
85%|████████▍ | 28/33 [00:01<00:00, 19.35it/s]
94%|█████████▍| 31/33 [00:01<00:00, 16.53it/s]
100%|██████████| 33/33 [00:01<00:00, 22.18it/s]
[<Axes: title={'center': 'Original'}, xlabel='kx axis (px)', ylabel='ky axis (px)'>, <Axes: title={'center': 'Difference of Gaussians'}, xlabel='kx axis (px)', ylabel='ky axis (px)'>, <Axes: title={'center': 'H-Dome'}, xlabel='kx axis (px)', ylabel='ky axis (px)'>]
Filtering Polar Images#
The available methods differ for Diffraction2D datasets and PolarDiffraction2D datasets.
Set the center of the diffraction pattern to its default, i.e. the middle of the image
s.calibration.center = None
Transform to polar coordinates
s_polar = s.get_azimuthal_integral2d(npt=100, mean=True)
s_polar_filtered = s_polar.subtract_diffraction_background(
"radial median",
inplace=False,
)
s_polar_filtered2 = s_polar.subtract_diffraction_background(
"radial percentile",
percentile=70,
inplace=False,
)
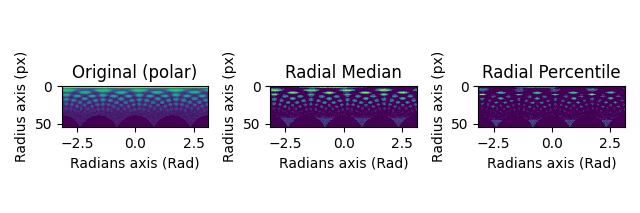
hs.plot.plot_images(
[s_polar.inav[2, 2], s_polar_filtered.inav[2, 2], s_polar_filtered2.inav[2, 2]],
label=["Original (polar)", "Radial Median", "Radial Percentile"],
tight_layout=True,
norm="symlog",
cmap="viridis",
colorbar=None,
)
0%| | 0/17 [00:00<?, ?it/s]
6%|▌ | 1/17 [00:03<00:54, 3.42s/it]
59%|█████▉ | 10/17 [00:03<00:01, 3.89it/s]
100%|██████████| 17/17 [00:03<00:00, 4.72it/s]
0%| | 0/33 [00:00<?, ?it/s]
100%|██████████| 33/33 [00:00<00:00, 404.10it/s]
0%| | 0/33 [00:00<?, ?it/s]
33%|███▎ | 11/33 [00:00<00:00, 78.10it/s]
58%|█████▊ | 19/33 [00:00<00:00, 71.62it/s]
82%|████████▏ | 27/33 [00:00<00:00, 62.80it/s]
100%|██████████| 33/33 [00:00<00:00, 71.35it/s]
[<Axes: title={'center': 'Original (polar)'}, xlabel='Radians axis (Rad)', ylabel='Radius axis (px)'>, <Axes: title={'center': 'Radial Median'}, xlabel='Radians axis (Rad)', ylabel='Radius axis (px)'>, <Axes: title={'center': 'Radial Percentile'}, xlabel='Radians axis (Rad)', ylabel='Radius axis (px)'>]
Total running time of the script: (0 minutes 11.650 seconds)
