simulated_strain#
- pyxem.data.simulated_strain(navigation_shape: tuple = (32, 32), signal_shape: tuple = (512, 512), disk_radius: int = 20, num_electrons: int = 100000.0, strain_matrix: ndarray[tuple[int, ...], dtype[_ScalarType_co]] = None, lazy: bool = False)[source]#
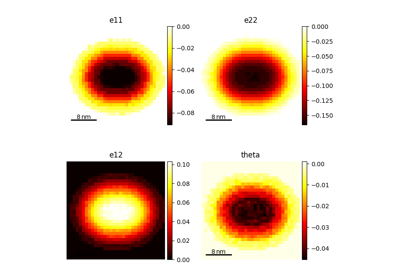
Create a simulated strain map from a simulated diffraction pattern and a strain matrix.
- Parameters:
navigation_shape (tuple) – The shape of the navigation axes, e.g. (32, 32).
signal_shape (tuple) – The shape of the signal axes, e.g. (512, 512).
disk_radius (int) – The radius of the disk used to create the diffraction pattern.
num_electrons – The number of electrons (per pixel) to simulate in the diffraction pattern.
strain_matrix – A 3x3 matrix representing the strain to apply to the diffraction pattern. If None, a default strain matrix is used.
lazy (bool) – If True, the returned signal will be lazy, otherwise it will be eager. Default is False.
- Returns:
A simulated diffraction pattern with applied strain.
- Return type: