Note
Go to the end to download the full example code.
Circular Hough Transform Peak Finding#
The Circular Hough Transform is a method to detect circular features in an image. It can be used to detect rings (or disks) in the diffraction pattern. For strain mapping there is some evidence that the Circular Hough transform is more accurate than typical template matching and can provide subpixel accuracy.
That being said, the Windowed Template Matching that pyxem uses is quite robust, (both to noise and in homogeniety in the diffraction disks) although a complete comparison hasn’t been fully studied.
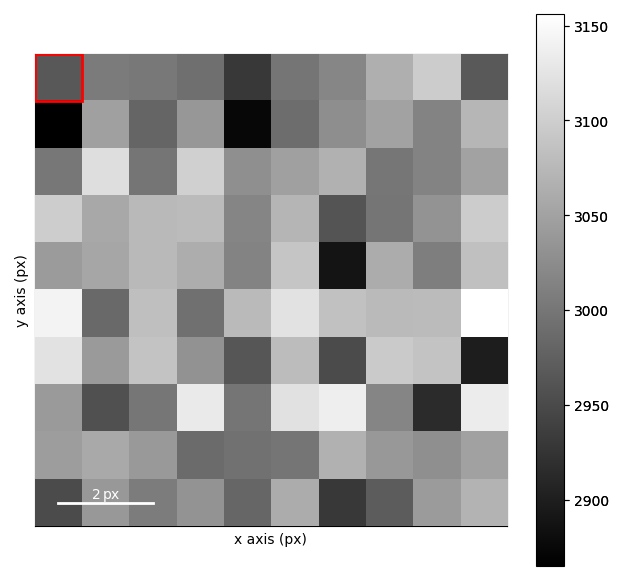
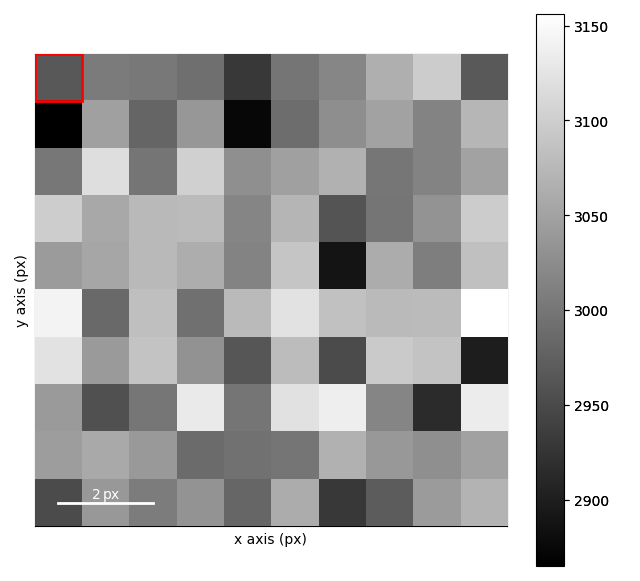
Making a Dummy Dataset
import hyperspy.api as hs
import pyxem as pxm
from skimage.transform import hough_circle
from skimage.feature import canny
import numpy as np
s = pxm.data.tilt_boundary_data(correct_pivot_point=True)
Canny Filter#
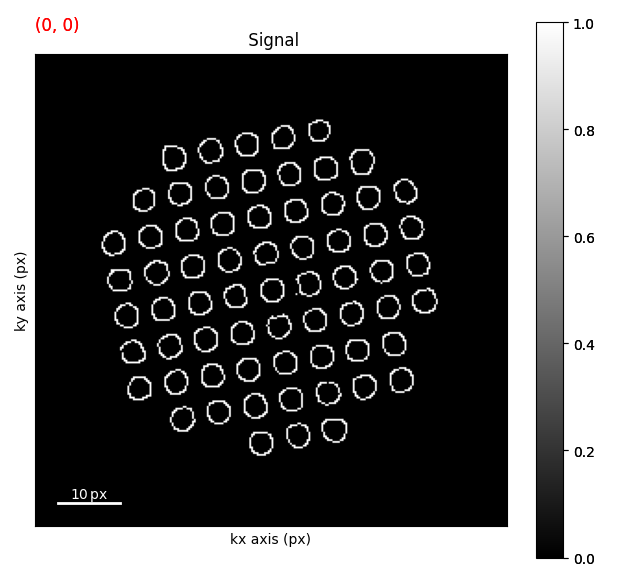
First we have to apply a Canny filter to the dataset to get a binary image of the outlines of the disks. This is basically a 1st diriviative in reciporical space
# Filter the image with a Canny filter.
canny_img = s.map(
canny,
sigma=2,
low_threshold=0.6,
high_threshold=0.8,
inplace=False,
use_quantiles=True,
)
canny_img.plot() # Plotting canny filtered image with outlines
0%| | 0/33 [00:00<?, ?it/s]
52%|█████▏ | 17/33 [00:00<00:00, 148.61it/s]
97%|█████████▋| 32/33 [00:00<00:00, 124.98it/s]
100%|██████████| 33/33 [00:00<00:00, 131.69it/s]
Computing the Hough Transform#
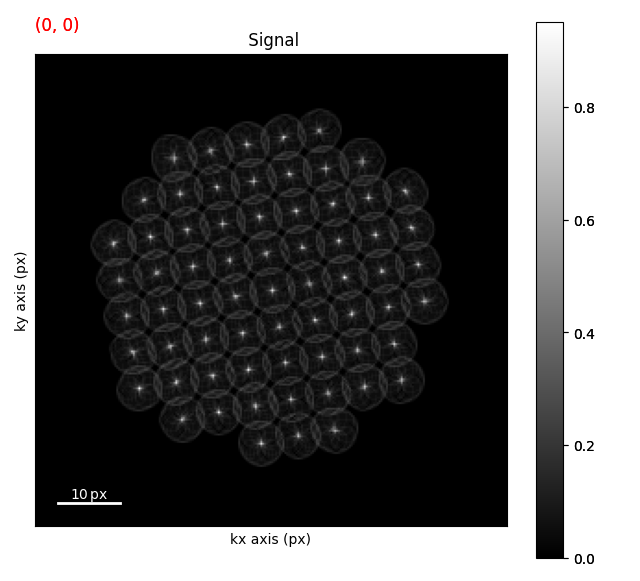
We can then compute the hough Transform using the radius of the disk. It is possible to have multiple radii but that will return multiple signals for each radii
def hough_circle_single_rad(img, radius, **kwargs):
return hough_circle(img, radius, **kwargs)[
0
] # Otherwise multiple radii are returned
circular_hough = canny_img.map(hough_circle_single_rad, radius=6, inplace=False)
circular_hough.plot()
0%| | 0/33 [00:00<?, ?it/s]
100%|██████████| 33/33 [00:00<00:00, 1046.47it/s]
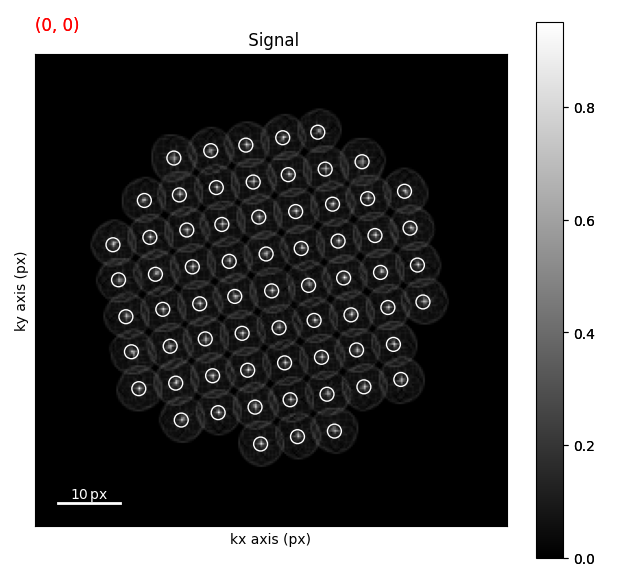
Finding Peaks#
Finding peaks is fairly easy from this point and uses the get_diffraction_vectors function.
dv = circular_hough.get_diffraction_vectors(threshold_abs=0.4, min_distance=4)
m = dv.to_markers(facecolor="none", edgecolor="w")
circular_hough.plot()
circular_hough.add_marker(m)
0%| | 0/33 [00:00<?, ?it/s]
64%|██████▎ | 21/33 [00:00<00:00, 207.74it/s]
100%|██████████| 33/33 [00:00<00:00, 215.72it/s]
0%| | 0/33 [00:00<?, ?it/s]
100%|██████████| 33/33 [00:00<00:00, 10992.06it/s]
0%| | 0/33 [00:00<?, ?it/s]
100%|██████████| 33/33 [00:00<00:00, 11450.37it/s]
sphinx_gallery_thumbnail_number = 6
Total running time of the script: (0 minutes 1.552 seconds)