Note
Go to the end to download the full example code.
Calibrating Scales#
This example demonstrates how to calibrate the scales in both reciprocal and real-space. Ideally this should be done using a calibration standard with a known lattice spacing.
There are many ways to calibrate the microscope so raise an issue if you have a specific method you would like to see implemented or if you have any questions.
from diffsims.generators.simulation_generator import SimulationGenerator
import pyxem as pxm
import hyperspy.api as hs
import numpy as np
from diffsims.utils.sim_utils import get_electron_wavelength
from scipy.optimize import curve_fit
import matplotlib.pyplot as plt
# Load the data and the cif file
au_dpeg = pxm.data.au_grating_20cm(
allow_download=True, signal_type="electron_diffraction"
)
gold_phase = pxm.data.au_phase(allow_download=True) # Orix.CrystalMap.Phase object
0%| | 0.00/263k [00:00<?, ?B/s]
5%|██▏ | 14.3k/263k [00:00<00:01, 136kB/s]
15%|█████▉ | 39.9k/263k [00:00<00:01, 199kB/s]
41%|████████████████▍ | 108k/263k [00:00<00:00, 405kB/s]
68%|███████████████████████████▏ | 178k/263k [00:00<00:00, 509kB/s]
0%| | 0.00/263k [00:00<?, ?B/s]
100%|███████████████████████████████████████| 263k/263k [00:00<00:00, 2.13GB/s]
0%| | 0.00/4.58k [00:00<?, ?B/s]
0%| | 0.00/4.58k [00:00<?, ?B/s]
100%|█████████████████████████████████████| 4.58k/4.58k [00:00<00:00, 43.8MB/s]
Create a Simulation Generator#
Use the SimulationGenerator to create a simulated diffraction pattern from the gold phase. Note that the acceleration voltage is irrelevant
sim_gen = SimulationGenerator()
sim1d = sim_gen.calculate_diffraction1d(gold_phase, reciprocal_radius=1.2)
Azimuthal Integration and Calibration#
The polar unwrapping for the 2D image will give a 1D diffraction pattern
that can be used for calibration. The calibration is done by comparing
the positions of the peaks in the simulated and experimental diffraction
patterns. Fitting with a hyperspy.model.Model1D can be used to find the
scale or the camera length.
au_dpeg.calibration.center = None # Set the center
az1d = au_dpeg.get_azimuthal_integral1d(npt=200, radial_range=(20, 128)) # in pixels
diffraction_model = az1d.model_simulation1d(sim1d, fit=True, center_lim=0.03)
theta_scale = az1d.model2theta_scale(
simulation=sim1d, model=diffraction_model, beam_energy=200
)
camera_length = az1d.model2camera_length(
simulation=sim1d,
model=diffraction_model,
beam_energy=200,
physical_pixel_size=5.5e-5,
)
print("Theta Scale: ", theta_scale, " Rad")
print(
"Camera Length: ", camera_length * 100, " cm"
) # Camera length of 21.287cm (vs 20cm from the microscope)
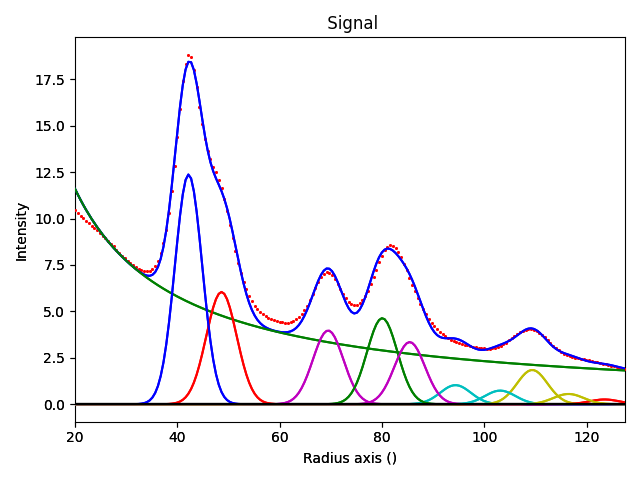
diffraction_model.plot(plot_components=True)
0%| | 0/2 [00:00<?, ?it/s]
100%|██████████| 2/2 [00:00<00:00, 1154.50it/s]
Theta Scale: 0.0002542800394167303 Rad
Camera Length: 21.6296957247437 cm
Aside: Showing how the Camera Length and scale is caluclated:#
centers = np.sort(
[
c.centre.value
for c in diffraction_model
if isinstance(c, hs.model.components1D.Gaussian)
]
)
wavelength = get_electron_wavelength(200)
angles = np.arctan2(
np.sort(sim1d.reciprocal_spacing), 1 / wavelength
) # both in inverse angstroms.
def f(x, m):
return x * m
m, pcov = curve_fit(f, centers, angles)
scale = m[0]
fig, axs = plt.subplots(1)
axs.scatter(centers, angles * 1000)
axs.plot(np.sort(centers), np.sort(centers) * scale * 1000)
axs.set_ylabel("Angle, mrad")
axs.set_xlabel("Detector Pixels")
axs.annotate(f"Slope: {scale*1000:.2} mrad/pixel", (0.1, 0.6), xycoords="axes fraction")
axs.annotate(
f"Camera Length: {5.5E-5/ np.tan(scale)*100:.4}cm",
(0.05, 0.5),
xycoords="axes fraction",
)
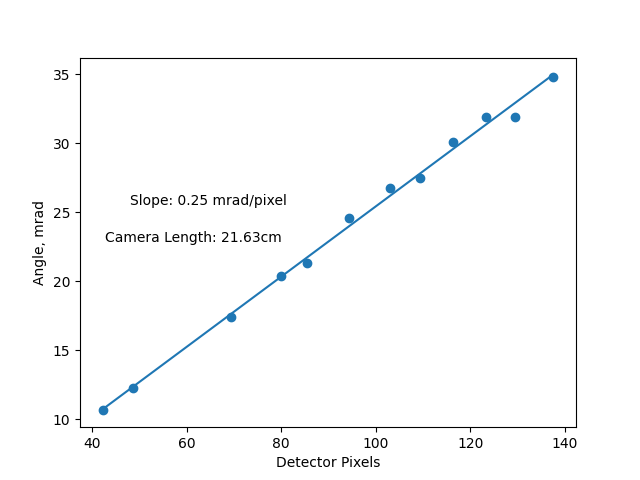
Text(0.05, 0.5, 'Camera Length: 21.63cm')
Applying a Camera Length#
The (calibrated) camera length can be applied to the data to calibrate the scale more accurately than a simple single point calibration. Let’s apply the camera length to the data and plot the result.
au_dpeg.calibration.detector(
pixel_size=5.5e-5, detector_distance=camera_length, beam_energy=200
)
print(au_dpeg.axes_manager) # non uniform axes
au_dpeg.plot(vmax="99th") # non uniform axes
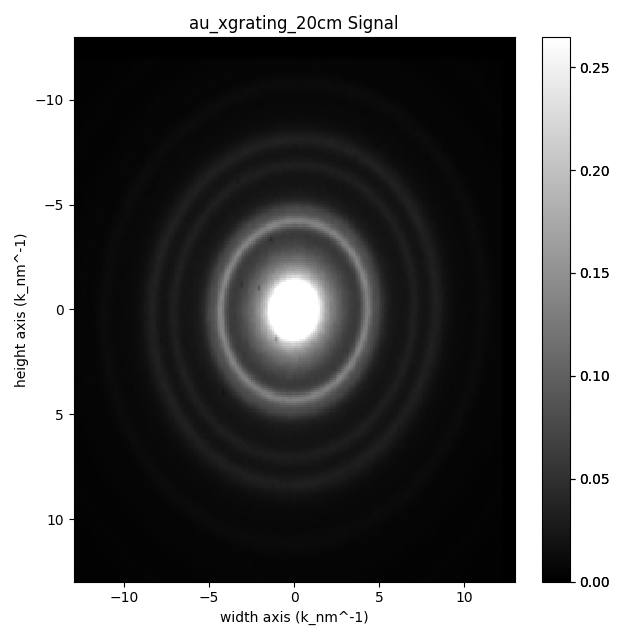
<Axes manager, axes: (|256, 256)>
Name | size | index | offset | scale | units
================ | ====== | ====== | ======= | ======= | ======
---------------- | ------ | ------ | ------- | ------- | ------
width | 256 | 0 | non-uniform axis | k_nm^-1
height | 256 | 0 | non-uniform axis | k_nm^-1
Plotting Corrected Azimuthal Integration#
When you get the Azimuthal Integration now that things are calibrated it automatically accounts for the ewald sphere now and uses the non uniform axes.
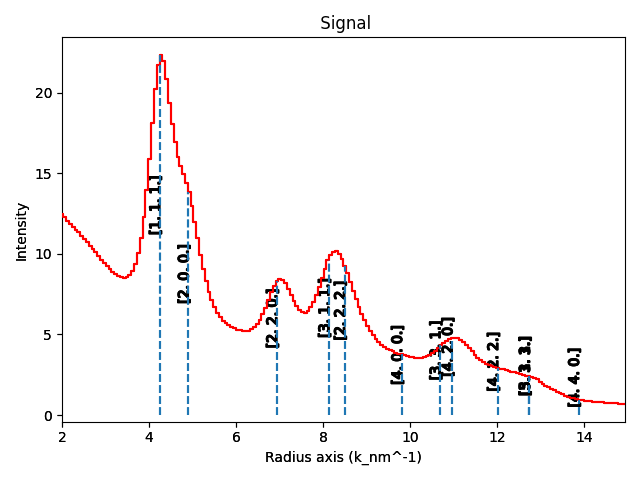
calibrated_azim = au_dpeg.get_azimuthal_integral1d(npt=200, radial_range=(2, 15))
calibrated_azim.plot()
x = np.array(sim1d.reciprocal_spacing) * 10
y = [
0.5,
] * len(x)
y0 = [
0,
] * len(x)
hkl = [str(h) for h in sim1d.hkl]
offsets = np.vstack((x, y)).T
t = hs.plot.markers.Texts(
offsets,
texts=hkl,
offset_transform="relative",
horizontalalignment="left",
verticalalignment="bottom",
rotation=np.pi / 2,
color="k",
)
calibrated_azim.add_marker(t)
lines = [[[l, 0], [l, 1]] for l in x]
v = hs.plot.markers.Lines(lines, transform="relative", linestyle="--")
calibrated_azim.add_marker(v)
0%| | 0/2 [00:00<?, ?it/s]
100%|██████████| 2/2 [00:00<00:00, 1357.38it/s]
Total running time of the script: (0 minutes 20.904 seconds)
